B.7.1 Describe the characteristics of biological catalysts (enzymes)
- enzymes increase the rate of reaction
- they can either decrease activation energy or improve chances of correct collision geometry, or both
- doesn't affect equilibrium position, change in enthalpy, or change in spontaneity
- not consumed or modified by the chemical reaction
- will eventually need to be replaced (generally a finite number of reactions or time before structure may degrade so that enzyme becomes non-functional)
- enzymes are proteins, complex, globular of 3° or sometimes 4° structure
- an enzyme as a catalyst is a particular for one specific reaction, named by what they do
B.7.2 Compare inorganic catalysts and biological catalysts (enzymes)
- most biological catalysts are protein molecules that undergo enzymatic reactions in aqueous solution in biological organisms
- e.g.) catalase
- inorganic catalysts can be surface metal catalysts where gas phase heterogenous reactions occur or they can be homogenous catalysts where the catalyst and reactant are in the same phase
- e.g.) heterogenous: nickel, homogenous: sulfuric acid in esterification reactions
- inorganic catalysts are not temperature sensitive while enzymes are because they're proteins and can be denatured at extreme temperatures beyond their optimal temperatures
B.7.3 Describe the relationship between substrate concentration and enzyme activity
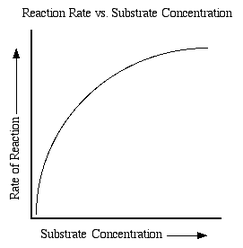
- if substrate concentration is increased (while other affecting factors such as pH and enzyme concentration are held constant), the rate of reaction increases and proceeds at a faster velocity
- this is because a greater substrate concentration means that there are a greater number of substrate particles to possibly hit and react with enzyme active sites per unit of time
- however, as the rate increases, the velocity levels off and slows down to reach Vmax, the maximum rate
- this is because all the active sites are engaged and there are no more free sites to react with substrate particles
- the only way to increase the rate further beyond this point is to increase enzyme concentration and thus add more free active sites
B.7.4 Determine Vmax and the value of the Michaelis-Menten constant (Km) by graphical means and explain its significance
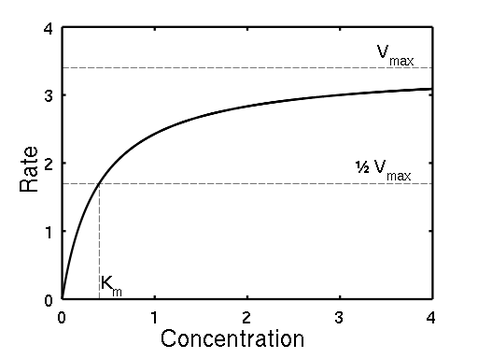
- Vmax of an enzyme is the max rate achieved when the enzyme concentration is kept constant and substrate concentration is increased
- Km (Michaelis constant) is the substrate concentration when the rate is 1/2 of the max rate
- Km can be used to compare the efficiency of different enzymes - the lower the Km the less amount of substrate is required to reach Vmax, so the enzyme is more efficient
- Vmax, 1/2 Vmax, and km can be found through a Michaelis-Menten plot, which has the substrate concentration as the x-values and the corresponding reaction rates as the y-values
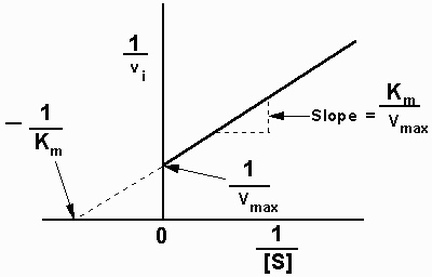
- A Lineweaver-Burke plot can also be used, but the x-values are instead 1/(substrate concentration) and the y-values are 1/(reaction rate)
B.7.5 Describe the mechanism of enzyme action, including enzyme substrate complex, active site, and induced fit model
How Enzymes Work:
Induced Fit Model:
What Hinders Enzyme Activity:
How Enzymes Work:
- the substrate (reactant) binds to a specific area on the enzyme called the active site
- the structure of the active site determines enzyme specificity, and can even be specific for an enantiomer
- visually the active site is a "pocket" or "groove" and is slightly deformable, able to change shape slightly to allow better binding of the substrate
Induced Fit Model:
- the enzyme and substrate are held together by relatively weak noncovalent interactions (hydrogen bonds, salt bridges, etc.)
- together this is called the enzyme-substrate complex
- the enzyme-substrate complex reacts and we get the enzyme-product complex
- the product then dissociates from the enzyme
- the free enzyme is then able to catalyze another reaction
What Hinders Enzyme Activity:
- enzymes can be irreversibly inactivated by pH extremely outside their optimal pH range and extreme temperature changes
- enzymes can also be reversibly inhibited
B.7.6 Compare competitive inhibition and non-competitive inhibition
Competitive Inhibition
Non-competitive Inhibition
Competitive Inhibition
- the inhibitor binds in the enzyme active site
- the inhibitor is very similar in structure to the substrate
- the substrate can't bind in the active site when the inhibitor is present
- individual enzyme molecules are completely inactivated until the inhibitor is removed
- overall the enzyme concentration is lowered and thus there's a slower reaction rate
- increasing substrate concentration can reduce competitive inhibitor effects
- e.g.) a substrate is succinate dehydrogenase and its inhibitor is malonate
Non-competitive Inhibition
- the inhibitor binds to the enzyme at a point other than the active site
- this causes some kind of conformational change in the active site that reduces enzyme effectiveness
- the inhibitor can either bind to the enzyme and prevent the substrate from binding or it can bind to the enzyme-substrate complex and prevent the reaction from proceeding
B.7.7 State and explain the effects of heavy-metal ions, temperature changes, and pH changes on enzyme activity
Temperature Change
pH Change
Heavy Metal Ions
Temperature Change
- increasing temperature will increase reaction rate as long as the temperature remains within the range of tolerance of the enzyme
- temperatures above the range will denature the enzyme
- critical temperature is the point of max reaction rate without denaturing
- e.g.) critical temperature is 37°C for humans
pH Change
- affects -NH2 and -COOH residues (small changes can be reversible)
- every enzyme has a particular optimum pH (e.g.) pH 2 for pepsin and pH 8 for chymotrypsin)
- different enzymes are more or less susceptible
- there is more impact if the affected residues are in the active site
Heavy Metal Ions
- ions of heavy metals such as lead, mercury, copper, or silver are non-competitive inhibitors for several enzymes
- e.g.) Silver ions react with -Sh groups in the side groups of cysteine residues in the protein chain. If the location of the cysteine residue is somewhere that affects tertiary structure, then the shape of the active site would be altered, preventing the enzyme from working